|
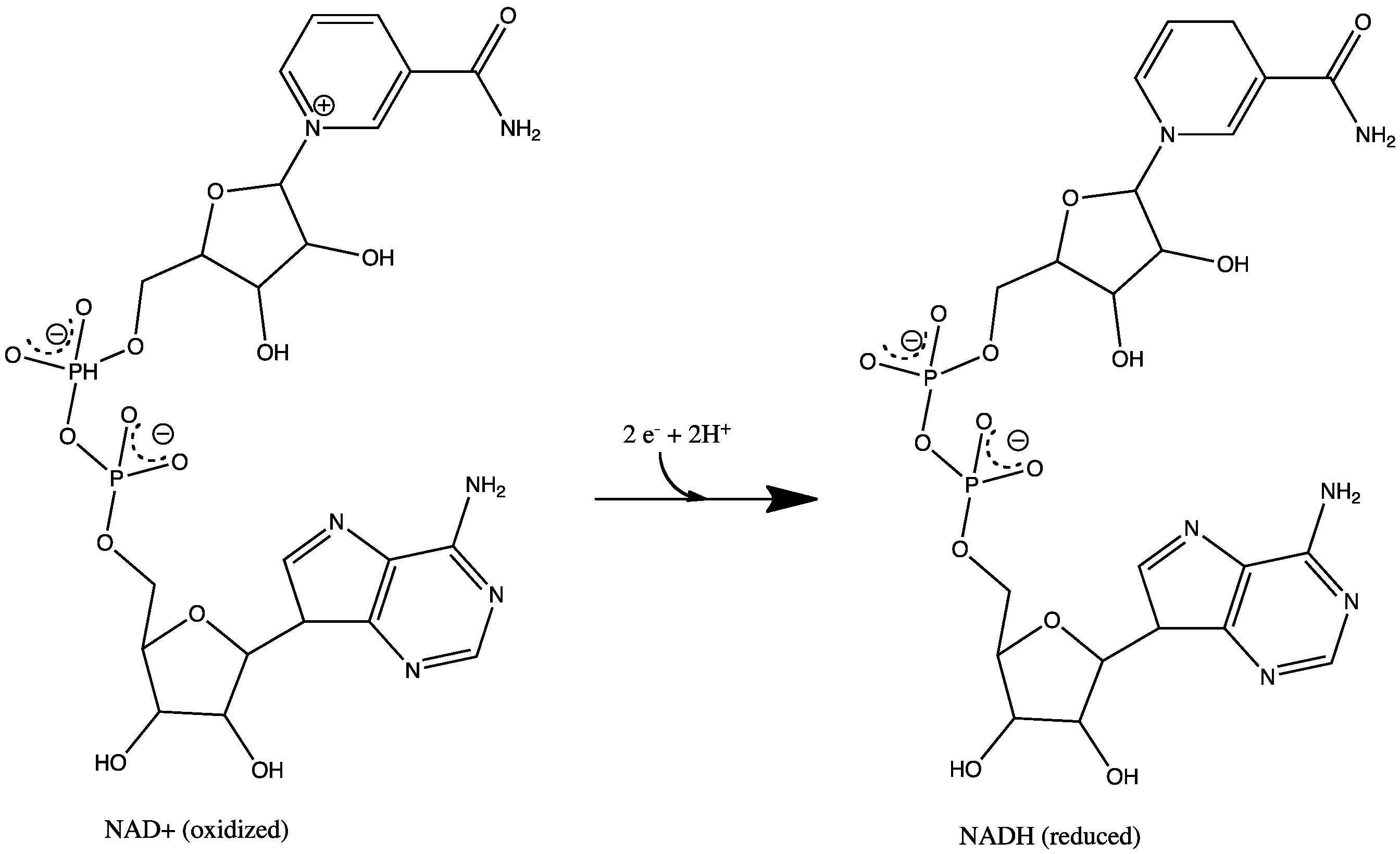
| NAD reduction |
Large collection of high quality biology pictures, photos, images, illustrations, diagrams and posters on marine biology, cell biology, microbiology... for educational purposes.
Electron Transport in Cellular Respiration
Cellular Respiration Pathway
 |
| Overview of Cellular Respiration |
Molecular Structure of the Chlorophyll
 |
| Molecular structure of Chlorophyll b |
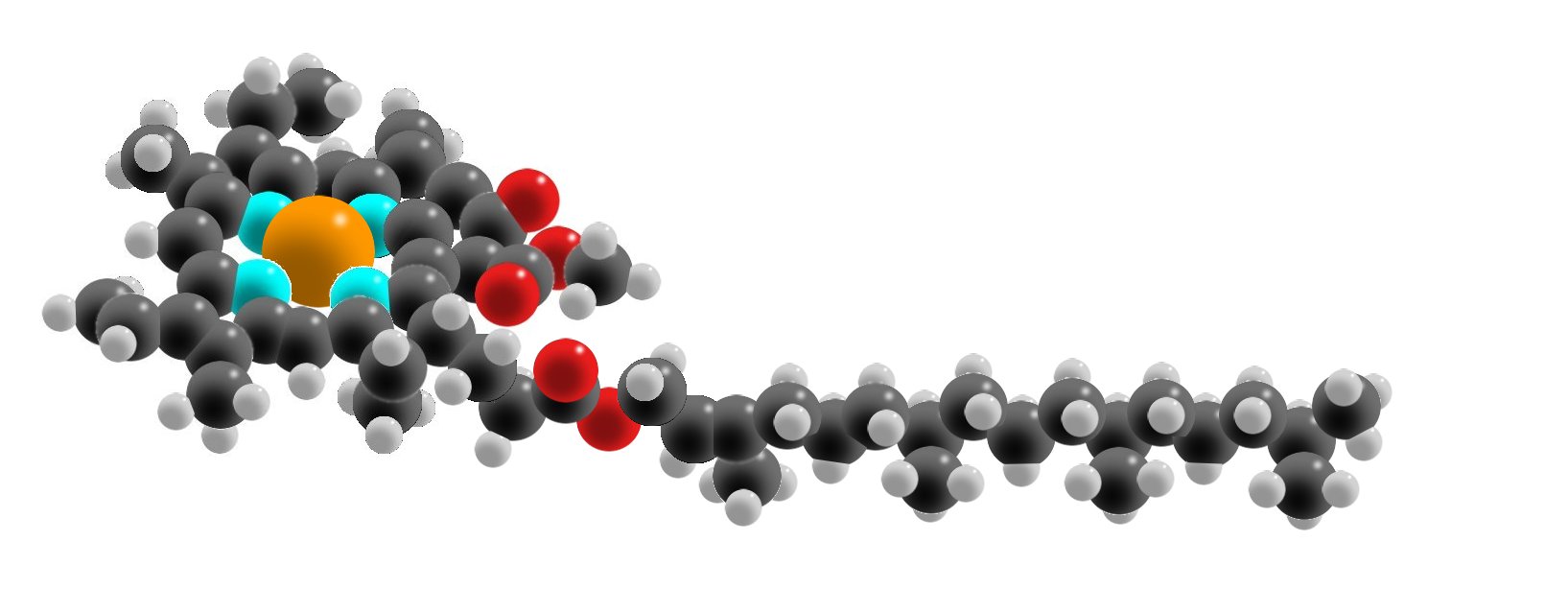
Chlorophyll is a green pigment absorbing light most strongly in the blue portion of the visible spectrum. This molecule is also known as a Photo-receptor. Chlorophyll has Carbon, Hydrogen, Oxygen, Nitrogen and Magnesium atoms. It has a Nitrogen ring at the top with a Magnesium atom center. This structure is also called Porphyrin Ring. As you see in the picture, porphyrin ring has alternating single and double bonds. Sidechains attached to the ring structure determines the type of the compound. The long hydrocarbon tail anchors the chlorophyll to the thylakoid membrane.
Space-filling Model of Chlorophyll
Brownish colored atoms at the left is Magnesium. 'Turquoise' colored balls are Nitrogen atoms. They hold the Magnesium atom and attaches it to porphyrin ring. Red colored balls represent Oxygen atoms. Hydrocarbon tail extending to the right has black colored Carbon atoms and white colored Hydrogen atoms.
Molecular Structure of FAD
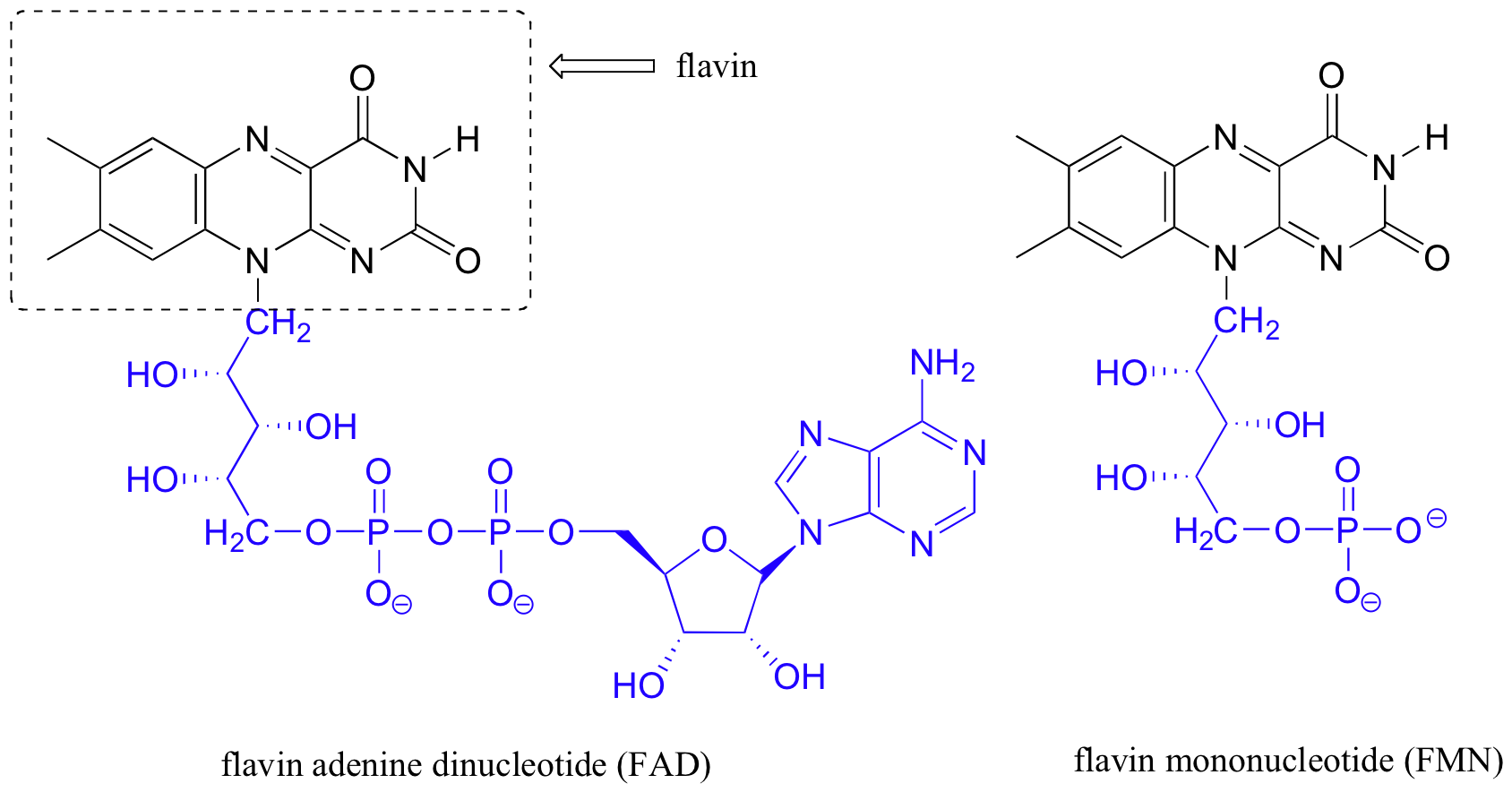 |
| FAD Structure |
Molecular Structure of NAD
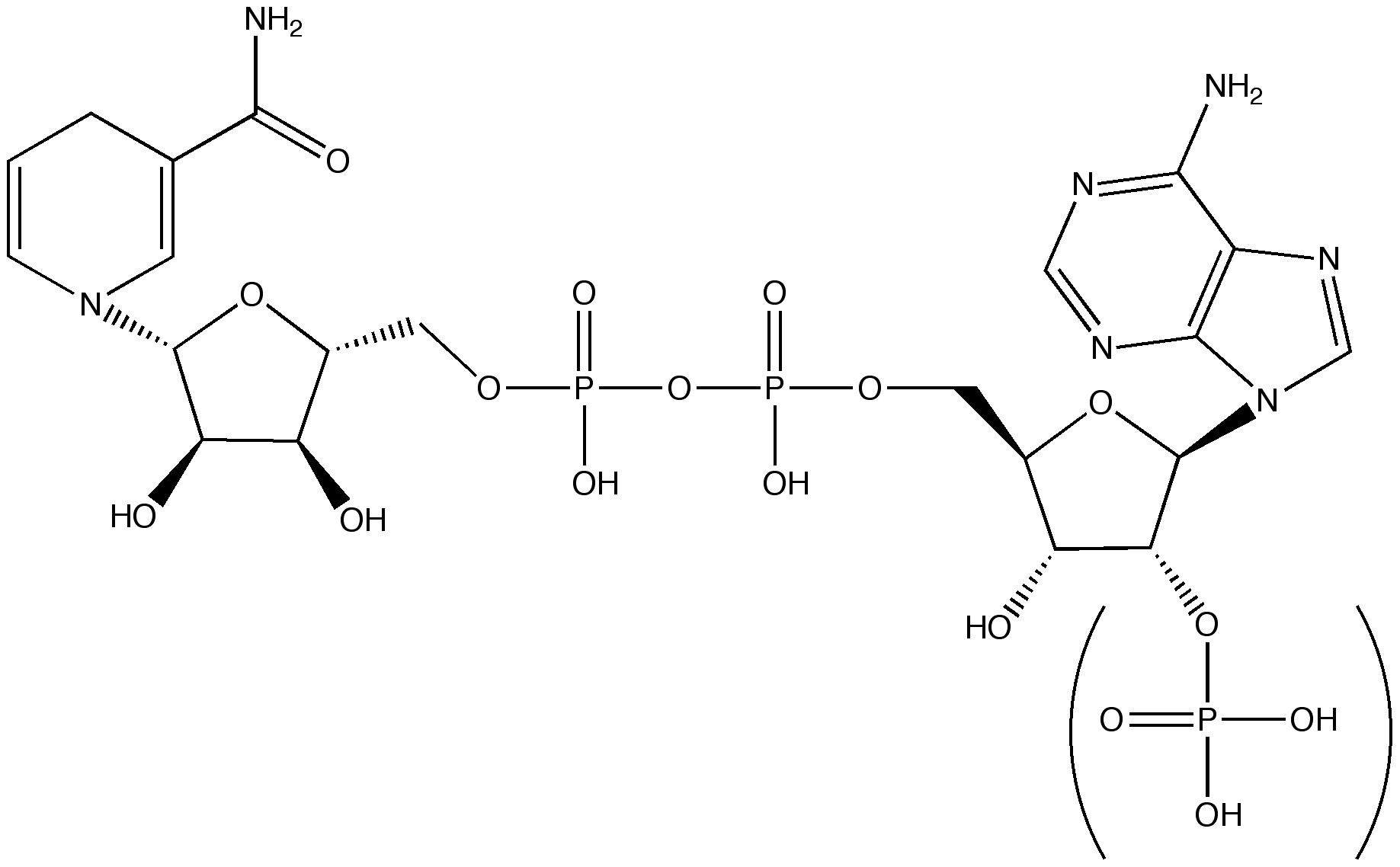 |
| Molecular Structure of NAD Molecule |
Sice they are coenzymes, some enzymes require them to work such as Alcohol Dehydrogenase, Pyruvate Dehydrogenase...
Molecular Structure of ATP
| Molecular Structure of ATP |
Table of Genetic Code
 |
| Genetic Code of Aminoacids |
Subscribe to:
Comments (Atom)